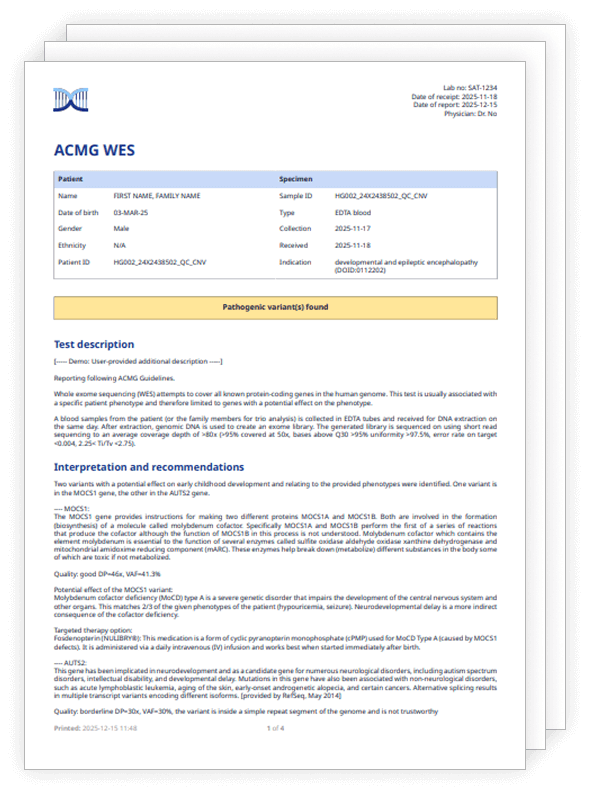
Why variant interpretation?
Flexible for you, from filtering strategies and virtual gene panels, to compatibility with any sequencer and kit. Whatever the clinical domain, variant interpretation puts the power in your hands.

Compliancy ensured
Generate genetic or oncology reports according to ACMG/AMP/ASCO/CAP guidelines. CE-IVD marked under IVDD (98/79/CE).

Quality integrated
Coverage analysis and other clinical QC metrics included automatically to reports. Define your own threshold values most important quality metrics.

All clinical domains
From clinical oncology to rare diseases diagnostics, variant interpretation streamlines and automates any clinical domain the lab expands into.

Safe and secure
Can be used from the cloud or installed on your premises behind the firewalls to keep confidential patient data safe and secured.

The tools are in your hands
With access to over 150 filterable metrics and integration of large amounts of data, you have everything you need in one place to get the most out of your data.
Flexible for you
Edit your filtering rules, report templates, virtual gene panels and much more from within the software. variant interpretation works for you and your needs.
Key features
Variant interpretation is built with flexibility as its core, to help you get the most from your NGS data
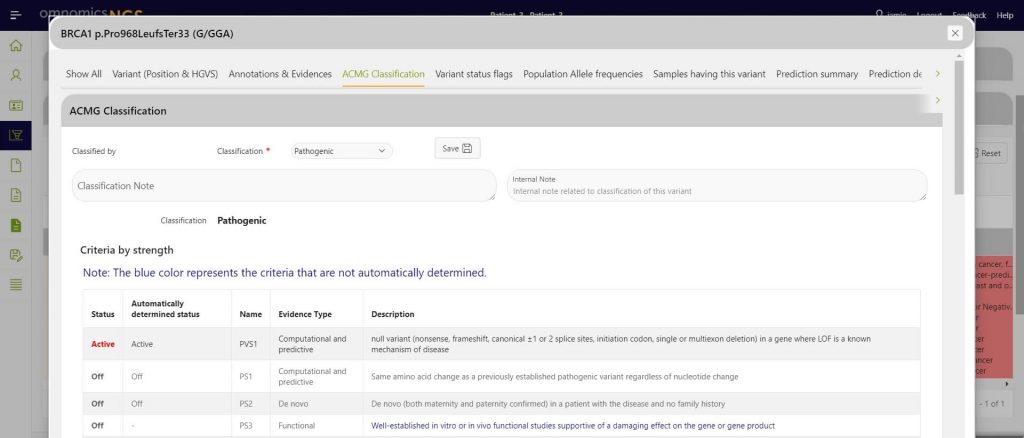
Automatic ACMG classification
Quickly and automatically classify variants based on the ACMG classification guidelines and manually change classification based on external knowledge gained through your own analyses.
Oncology, rare diseases, carrier screening
Regardless of your clinical domain, Variant interpretation works for you. Run tumour/normal samples, dig deep into WES or WGS data to uncover variants causing rare diseases, run dual sample carrier screening for deleterious variants or trace hereditary diseases through duo, trio and family analyses.

Database integration
One of the most extensive database integrations available. All the data you need at your fingertips. ClinVar for germline, CIViC for cancer, HPO, OMIM, Ensembl, OrphaNet, CGC, Reactome and more. Frequently updated and a list that is always expanding.

Flexible filtering
Quickly build filtering strategies based on 150 filterable data columns. Make your analyses reliable and reproducible by implementing the same filtering strategy every time, and dig deep into large samples. Further filter by enacting customisable virtual gene panels.

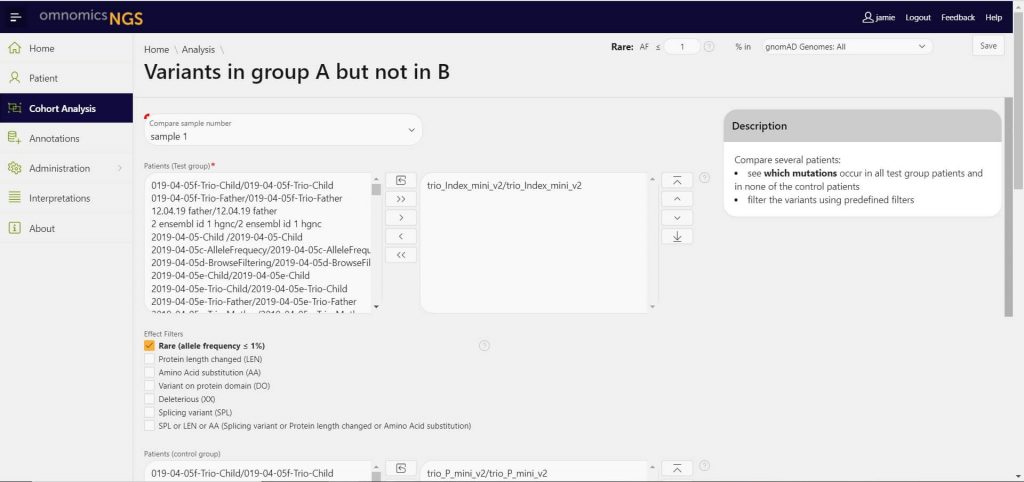
Cohort analysis
Easily group samples based on cohorts to understand potential commonality between samples. Quickly highlight shared variants, and enact filters to highlight only those relevant variants.
Pipeline integration
Variant interpretation is designed to easily integrate into whatever pipeline you already have, including electronic patient record systems, LIMS and more. Let Variant interpretation become a seamless part of your operations.

CNV Analysis
The CNV workbench has been designed to greatly reduce the effort involved in CNV classification by comprehensively calculating the ACMG/ClinGen classification score and allowing to visualise the result in compact as well as fully documented mode.

Digging deep into WGS samples for studying autism spectrum disorders
Variant interpretation is being used to study WGS samples from cohorts of patients around the world as part of the GEMMA project to study autism spectrum disorders. The capabilities of the tool allow us to explore the hundreds of thousands of variants in each sample, highlighting those which may play a role in explaining ASD. Versatile as both a research and clinical tool, variant interpretation gives you the power to dig deep into any sample.






